Write internal xspec data to a tcl variable. This facility allows the
manipulation of xspec data by tcl scripts, so that one can, for example,
extract data from xspec runs and store in output files, format xspec
output data as desired, use independent plotting software, etc.
| ? |
Show the valid options. Does not set $xspec_tclout. |
| areascal n <s|b> |
Writes a string of blank separated values giving the AREASCAL values for spectrum n.
If no second argument is given or it is “s” then the values are from the source file, if “b” from the background file. |
| arf n |
The auxiliary response filename(s) for spectrum n. |
| backgrnd n |
Background filename for spectrum n. |
| backscal n <s|b> |
Same as areascal option but for BACKSCAL value. |
| chain best|dic|last|proposal|
stat |
The best option returns
the parameter values corresponding to the smallest statistic value in the
loaded chains. The dic option returns the deviance information
criterion and effective number of parameters calculated by the last
chain dic command. The last option returns the final set of parameter values in
the loaded chains. The proposal option takes arguments distribution or matrix
and returns the name or covariance matrix for the proposal distribution when
using Metropolis-Hastings. The stat option returns the output of the
last chain stat command. |
| chatter |
Current xspec chatter level. |
| compinfo [<mod>:]n [<group n>] |
Name, 1st parameter number and number
of parameters of model component n, belonging to model w/ optional name <mod>
and optional datagroup <group n>. |
| cosmo |
Writes a blank separated string containing the Hubble constant (H0),
the deceleration parameter (q0), and the cosmological constant (Lambda0).
Note that if Lambda0 is non-zero the Universe is assumed to be flat and
the value of q0 should be ignored. |
| covariance [m,n] |
Element (m,n) from the covariance matrix of the most recent fit.
If no indices are specified, then entire covariance matrix is retrieved. |
| datagrp [n] |
Data group number for spectrum n. If no n is given, outputs the
total number of data groups. |
| datasets |
Number of datasets. |
| dof |
Degrees of freedom in fit, and the number of channels. |
| energies [n] |
Writes a string of blank separated values giving the
energies for spectrum n on which the model is calculated. If n is not
specified or is 0, it will output the energies of the default dummy
response matrix. |
| eqwidth n [errsims] |
Last equivalent width calculated for spectrum n.
If errsims keyword is supplied, this will instead return the
complete sorted array of values generated for the most recent eqwidth error
simulation. |
| error [<mod>:]n (for gain parameters use: rerror [<sourceNum>:]n) |
Writes last confidence region calculated for parameter n of model with optional
name <mod>, and a string listing any errors that occurred during the
calculation. The string comprises nine letters, the letter is T or F
depending on whether or not an error occurred. The 9 possible errors are:
- new minimum found
- non-monotonicity detected
- minimization may have run into problem
- hit hard lower limit
- hit hard upper limit
- parameter was frozen
- search failed in -ve direction
- search failed in +ve direction
- reduced chi-squared too high
So for example an error string of “FFFFFFFFT” indicates the calculation
failed because the reduced chi-squared was too high. |
| expos n <s|b> |
Same as areascal option but for EXPOSURE value. |
| filename n |
Filename corresponding to spectrum n. If the
filename is type II then include the {n} suffix specifying the row
from which the spectrum was read. |
| fileinfo <keyword> n |
Value of keyword in the SPECTRUM
extension of the file corresponding to spectrum n. |
| flux [n][errsims] |
Last model flux or luminosity calculated for spectrum n.
Writes a string of 6 values: val errLow errHigh (in ergs/cm2) val errLow
errHigh (in photons). Error values are .0 if flux was not run with errsims option.
If the errsims keyword is supplied, this will instead return the
completed sorted array of values generated during the most recent flux error calculation. |
| ftest |
The result of the last ftest command. |
| gain [<sourceNum>:]<specNum>slope|
offset |
For gain fit parameters: value, delta, min, low, high, max for the slope or offset
parameter belonging to the [<sourceNum>:]<specNum>response.
For nonfit gain parameters, only the value is returned. |
| goodness [sims] |
The percentage of realizations from the last goodness
command with statistic value less than the best-fit statistic using the data.
If optional sims keyword is specified, this will instead give the
full array of simulation values from the last goodness command. |
| idline e d |
Possible line IDs within the range [e-d, e+d]. |
| ignore [<n>] |
The range(s) of the ignored channels for spectrum n. |
| lumin [n] [errsims] |
Last model luminosity calculated for spectrum n.
Same output format as flux option, in units of
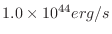 . . |
| margin probability|fraction|
[<modName>:]<parNum> |
The probability and fraction options return the probability and
fraction columns, respectively, from the
most recent margin command. Otherwise, the parameter column indicated by
<parNum>is returned. Note that for multi-dimensional margin the returned
parameter column will contain duplicate values, in the same order as they
originally appeared on the screen during the margin run. |
| model |
Description of current model(s). |
| modcomp [<modName>] |
Number of components in model (with optional model name). |
| modgroups [<modName>] |
The data group numbers associated with a
model (with optional model name). |
| modnames |
The names of models. |
| modkeyval [<key>] |
The value attached to <key>in
the model (key,value) database. Key and value pairs can be added to
this database by calling in a function routine the method
loadDbValue(const string key, Real value) for C++ or
PDBVAL(character*(*) key, double precision value) for Fortran. A list
of the current (key,value) pairs can be obtained by entering “?” for
<key>. |
| modpar [<modName>] |
Number of model parameters(with optional model name). |
| modval [<specNum>[<modName>]] |
Write to Tcl the last calculated model
values for the specified spectrum and optional model name. Writes a string of
blank separated numbers. Note that the output is in units of
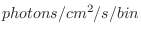 . . |
| nchan [<n>] |
Total number of channels in spectrum n (including ignored channels). |
| noticed [<n>] |
Range (low,high) of noticed channels for spectrum n. |
| noticed energy [<n>] |
The noticed energies for spectrum n. |
| nullhyp |
When using chi-square for fits, this will retrieve the reported null
hypothesis probability. |
| param [<mod>:]n |
(value, delta, min, low, high, max) for model parameter n. |
| peakrsid n [lo,hi] |
Energies and strengths of the peak residuals (+ve and -ve)
for the spectrum n. Optional arguments lo, hi specify an energy range in
which to search. |
| pfree [<mod>:]n |
T returned if parameter is free, F if not. |
| pinfo [<mod>:]n |
Parameter name and unit for parameter n of model
with optional name. |
| plink [<mod>:]n |
Information on parameter linking for parameter n.
This is in the form true/false (T or F) for linked/not linked, followed
by the multiplicative factor and additive constants if linked. |
| plot <option><array>[<plot group n>] |
Write a string of blank separated values for the array. <option>
is one of the valid arguments for the plot or iplot commands. <array>
is one of x, xerr, y, yerr, or model. xerr and yerr output the 1-sigma error
bars generated for plots with errors. The model array is for the convolved
model in data and ldata plots. For contour plots this command just dumps the
steppar results. The command does not work for genetic plot options. |
| plotgrp |
Number of plot groups. |
| query |
The setting of the query option. |
| rate <n|all> |
Count rate, uncertainty, model rate, and
percentage of net flux (no background) compared to total flux for the specified
spectrum n, or for the sum over all spectra. |
| rerror [<sourceNumber>:]n |
Writes last confidence region calculated for response
parameter n of model with optional source number, and a string listing any
errors that occurred during the calculation. See the help above on the
error option for a description of the string. |
| response n |
Response filname(s) for the spectrum n. |
| sigma [<modelName>:]n |
The sigma uncertainty value for parameter n.
If n is not a variable parameter or fit was unable to calculate sigma, -1.0 is returned. |
| simpars |
Creates a list of parameter values by drawing from a multivariate
Normal distribution based on the covariance matrix from the last fit.
This is the same mechanism that is used to get the errors on fluxes and
luminosities, and to run the goodness command. |
| solab |
Solar abundance table values. |
| stat [test|<n>] |
Value of statistic. If optional test argument is given,
this outputs the test statistic rather than the fit statistic. If a spectrum number n is given, the fit statistic contribution from that one spectrum
is returned. |
| statmethod [test] |
The name of the fit stat method currently in use.
If optional test argument is given, this will give the name of the
test stat method. |
| steppar statistic | delstat |
[<modName>:]<parNum> |
The statistic and delstat options return the statistic or
delta-statistic column respectively from the most recent steppar run.
Otherwise, the parameter column indicated by <parNum>is returned.
Note that for multi-dimensional steppars the returned parameter column will
contain duplicate values, in the same order as they originally appeared on
the screen during the steppar run. |
| systematic [<modName>|unnamed] |
The current model systematic error values.
If no argument is given, this returns the default systematic value applied to
all models except those that have explicitly been assigned their own value. If
a <modName>is given, this returns the systematic error value for that
specific model. This will be the same as the default setting unless it has
been explicitly assigned its own value. To get the specific setting for an unnamed
model, enter unnamed. |
| varpar |
Number of variable fit parameters. |
| version |
The XSPEC version string. |
| weight |
Name of the current weighting function. |
| xflt n |
XFLT#### keywords for spectrum n. The first number written is the
number of keywords and the rest are the keyword values.
|
![]() statistic and the three
parameters to tcl variables $chistat, $par1, $par2, and $par3. These can
now be manipulated in any way permitted by tcl. Examples of using tclout and
tcloutr can be found in the Xspec/src/scripts directory.
statistic and the three
parameters to tcl variables $chistat, $par1, $par2, and $par3. These can
now be manipulated in any way permitted by tcl. Examples of using tclout and
tcloutr can be found in the Xspec/src/scripts directory.