THE XMM-NEWTON ABC GUIDE, STREAMLINED
EPIC (IMAGING mode), Hera Command Window
Contents
Prepare the Data
Reprocess the Data
Make a Light Curve
Apply Standard Filters
Apply Time Filters
Source Detection
Make an Exposure Map
Extract the Source and Background Spectra
Determine the Spectrum Extraction Areas
Check for Pile Up
My Observation is Piled Up! Now What?
Create the Photon Redistribution Matrix (RMF) and Ancillary File (ARF)
Prepare the Data
Please note that the two tasks in this section (cifbuild and odfingest) must be run in the ODF directory. These are the only tasks with that requirement, and after this section, we will work exclusively in our reprocessing directory.Many SAS tasks require calibration information from the Calibration Access Layer (CAL). Relevant files are accessed from the set of Current Calibration File (CCF) data using a CCF Index File (CIF). Setting the environment paraters follows the same syntax as the cshell in linux. To set the environment parameters and make the ccf.cif file, navigate into the ODF directory and in the Command Window, type
setenv SAS_ODF /data/0123700101/ODF setenv SAS_ODFPATH /data/0123700101/ODF cifbuildTo use the updated CIF file in further processing, you will need to reset the environment variable SAS_CCF:
setenv SAS_CCF /data/0123700101/ODF/ccf.cifThe task odfingest extends the Observation Data File (ODF) summary file with data extracted from the instrument housekeeping data files and the calibration database. It is only necessary to run it once on any dataset, and will cause problems if it is run a second time. If for some reason odfingest must be rerun, you must first delete the earlier file it produced. This file largely follows the standard XMM naming convention, but has SUM.SAS appended to it. To run odfingest and reset the environment variable:
odfingest setenv SAS_ODF /data/0123700101/ODF/0070_0123700101_SCX00000SUM.SAS
Reprocess the Data
To reprocess the data, go up one directory in the tree and make a new working directory, either using the buttons in the upper left corner of the User Account Window or the commands you would expect with a linux machine, as seen below. When you are in the working directory, call either emchain or emproc for MOS data, and epchain or epproc for PN data.cd .. mkdir reproc cd reproc emchainor
emprocand
epchainor
epproc
By default, none of these tasks keep any intermediate files they generate. Emchain and epchain maintain the usual naming convention. Emproc and epproc designate their output event files with "ImagingEvts.ds". In any case, it is convenient to rename them something easy to type; this is mostly easily done by clicking on the pen icon next to the file name in the User Account Window. We'll assume the new name for the MOS1 event file is mos1.fits.
Make a Light Curve
The XMM-Newton Observatory is susceptible to soft particle flaring, so it is necessary to examine the light curve to determine how much of the data is useful.
To create a light curve, type
evselect table=mos1.fits rateset=mos1_ltcrv.fits maketimecolumn=yes
timebinsize=100 makeratecolumn=yes
where
-
table - input event table
rateset - name of output light curve file
maketimecolumn - make a time column
timebinsize - time binning (seconds)
makeratecolumn - make a count rate column, otherwise a count column will be created
The output file mos1_ltcrv.fits can be viewed by downloading and displaying it with fv.
fv mos1_ltcrv.fits &
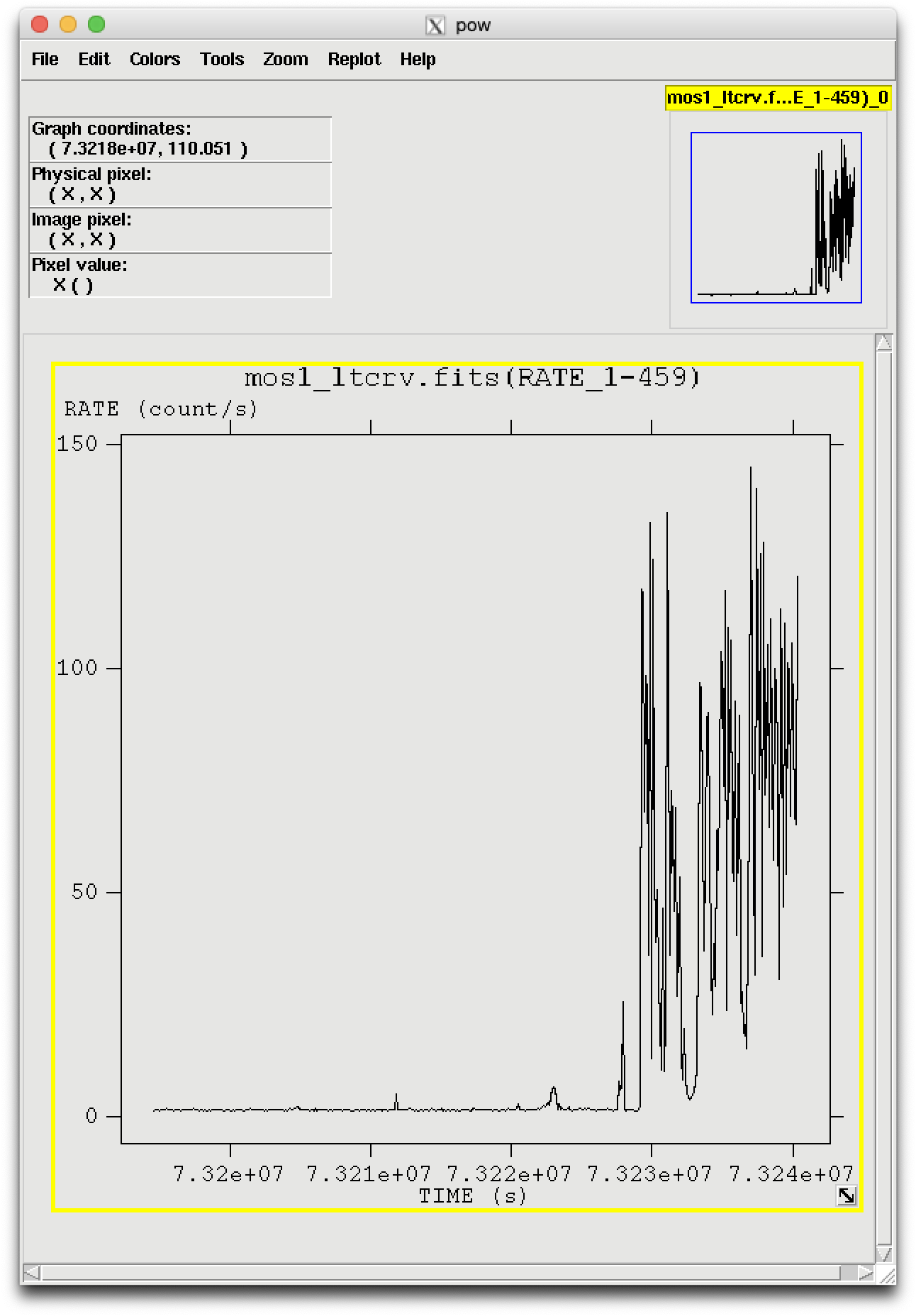
In the pop-up window, the RATE extension will be available in the second row (index 1, as numbering begins with 0). Selecting PLOT from this row will let you choose the column name and axis on which to plot it. The light curve is shown in Figure 1.
Apply Standard Filters
The filtering expressions for the MOS and PN are:
(PATTERN <= 12)&&(PI in [200:12000])&&#XMMEA_EM
and
(PATTERN <= 12)&&(PI in [200:15000])&&#XMMEA_EP
If you are going to extract a spectrum from the PN data, you will need the most stringent filters in order to get a high-quality spectrum:
(PATTERN <= 4)&&(PI in [200:15000])&&#XMMEA_EP&&(FLAG == 0)
For the MOS, the standard filters should be appropriate for most cases.
The first two expressions will select good events with PATTERN in the 0 to 12 range. The PATTERN value is similar the GRADE selection for ASCA data, and is related to the number and pattern of the CCD pixels triggered for a given event.The PATTERN assignments are: single pixel events: PATTERN == 0, double pixel events: PATTERN in [1:4], triple and quadruple events: PATTERN in [5:12].
The second keyword in the expressions, PI, selects the preferred pulse height of the event; for the MOS, this should be between 200 and 12000 eV. For the PN, this should be between 200 and 15000 eV. This should clean up the image significantly with most of the rest of the obvious contamination due to low pulse height events. Setting the lower PI channel limit somewhat higher (e.g., to 300 eV) will eliminate much of the rest.
Finally, the #XMMEA_EM (#XMMEA_EP for the PN) filter provides a canned screening set of FLAG values for the event. (The FLAG value provides a bit encoding of various event conditions, e.g., near hot pixels or outside of the field of view.) Setting FLAG == 0 in the selection expression provides the most conservative screening criteria and should always be used when serious spectral analysis is to be done on the PN. It typically is not necessary for the MOS.
To filter the data, type
evselect table=mos1.fits filtertype=expression filteredset=mos1_filt.fits
expression='(PATTERN <= 12) && (PI in [200:12000]) && #XMMEA_EM'
where
-
table - input event table
filtertype - method of filtering
expression - filtering expression
filteredset - output file name
Apply Time Filters
Sometimes, it is necessary to use filters on time in addition to those mentioned above. This is because of soft proton background flaring, which can have count rates of 100 counts/sec or higher.
It should be noted that the amount of flaring that needs to be removed depends in
part on the object observed; a faint, extended object will be more affected than
a very bright X-ray source.
There are two ways to filter on time: with an explicit reference to the TIME parameter
in the filtering expression, or by creating a secondary Good Time Interval
(GTI) file with the task tabgtigen. Both procedures are described below. For the
example data, we will filter by time, though you can just as easily filter by rate.
To explicitly define the TIME parameter, make a light curve and display it, as demonstrated above.
There is a very large flare toward the end of the observation, so the syntax for the time selection is (TIME <= 73227600). However, there is also a small flare within an otherwise good interval. A slightly more comlicated expression to remove it would be: (TIME <= 73227600)&&!(TIME IN [73221920:73223800]). The syntax &&(TIME < 73227600) includes only events with times less than 73227600, and the "!" symbol stands for the logical "not", so use &&!(TIME in [73221920:73223800]) to exclude events in the time interval 73221920 to 73223800.
If combined with the standard filtering expression, the full filtering expression would then be:
(PATTERN <= 12) && (PI in [200:12000]) && #XMMEA_EM &&
(TIME <= 73227600) &&! (TIME in [73221920:73223800])
This expression can then be used to
filter the original event file, or only the times can be used to filter the
file that has already had the standard filters applied:
evselect table=mos1_filt.fits filtertype=expression filteredset=mos1_filt_time.fits
expression='(TIME <= 73227600) &&!(TIME in [73221920:73223800])'
where the keywords are as described above.
To filter on time using a secondary GTI file, make the file by using the same time filtering parameters as determined above and the tabgtigen task,
tabgtigen table=mos1_ltcrv.fits gtiset=gtiset.fits timecolumn=TIME
expression='(TIME <= 73227600)&&!(TIME in [73221920:73223800])'
where
-
table - input count rate table
expression - filtering expression
gtiset - output file name for selected GTI intervals
timecolumn - time column
and apply the new GTI file with evselect:
evselect table=mos1_filt.fits filtertype=expression filteredset=mos1_filt_time.fits
expression='GTI(gtiset.fits,TIME)'
where the evselect parameters are as described previously. The event file can be viewed by downloading it and viewing it locally with ds9. An image of the MOS1 events file with both the standard and time filters applied is shown in Figure 2.
Source Detection
The edetect_chain task does nearly all the work involved with EPIC source detection. It can process up to three intruments (both MOS cameras and the PN) with up to five images in different energy bands simultaneously. All images must have identical binning and WCS keywords. For this example, we will perform source detection on MOS1 images in two bands (``soft'' X-rays with energies between 300 and 2000 eV, and ``hard'' X-rays, with energies between 2000 and 10000 eV) using the filtered event files produced here. Edetect_chain has many parameters that users might want to edit, such as detection likelihood threshold; more information about them can be found here.
We will start by generating some files that edetect_chain needs: an attitude file and images of the sources in the desired energy bands, with the image binning sizes as needed according to the detector. For the MOS, we'll let the binsize be 22.
First, the attitude file:
- atthkgen atthkset=attitude.fits
where
- atthkset - output file name
Next, the soft and hard X-ray images:
-
evselect table=mos1_filt_time.fits withimageset=yes imageset=mos1-s.fits
imagebinning=binSize xcolumn=X ximagebinsize=22 ycolumn=Y yimagebinsize=22
filtertype=expression expression='(FLAG == 0)&&(PI in [300:2000])'
and
-
evselect table=mos1_filt_time.fits withimageset=yes imageset=mos1-h.fits
imagebinning=binSize xcolumn=X ximagebinsize=22 ycolumn=Y yimagebinsize=22
filtertype=expression expression='(FLAG == 0)&&(PI in [2000:10000])'
where the parameters are as listed previously,
and
-
withimageset - flag to create an image
imageset - fits image name to be created
imagebinning - how to bin the image
xcolumn - table column to use for the X axis
ximagebinsize - binning in X axis
ycolumn - table column to use for the Y axis
yimagebinsize - binning in Y axis
Now we can run edetect_chain.
-
edetect_chain imagesets='mos1-s.fits mos1-h.fits' eventsets='mos1_filt_time.fits'
attitudeset=attitude.fits pimin='300 2000' pimax='2000 10000'
ecf='0.878 0.220'
where
-
imagesets - list of count images
eventsets - list of event files
attitudeset - attitude file name
pimin - list of minimum PI channels for the bands
pimax - list of maximum PI channels for the bands
ecf - energy conversion factors for the bands
The energy conversion factors (ECFs) convert the source count rates into fluxes. The ECFs for each detector and energy band depend on the pattern selection and filter used during the observation. For more information, please consult the 3XMM Catalogue User Guide. Those used here are derived from PIMMS using the flux in the 0.1-10.0 keV band, a source power-law index of 1.9, an absorption of 0.5×1020.
The task srcdisplay can be used to produce a region file for the sources (ignore the error messages.) This can be downloaded and overlayed on the event file with ds9.
srcdisplay boxlistset=emllist.fits imageset=mos1-s.fits
regionfile=regionfile.txt sourceradius=0.01 withregionfile=yes
where
-
boxlistset - eboxdetect source list
imageset - image file name over which the source circles are to be plotted
regionfile - file name of output file containing source regions
sourceradius - radius of circle plotted to locate sources
withregionfile - flag to create a region file
Figure 3 shows the MOS1 image overlayed with the detected sources.
Make an Exposure Map
Prior to making an exposure map, you will need to make an image in the energy band you are interested in. You will also need the attitude file from atthkgen. Let's say that we're interested in the 1-2 keV band; we can use the attitude file that we made before, and to make the image,
evselect table=mos1_filt_time.fits withimageset=yes imageset=mos1_1-2keV.fits
imagebinning=binSize xcolumn=X ximagebinsize=22 ycolumn=Y yimagebinsize=22
filtertype=expression expression='(PI in [1000:2000])'
Then,
eexpmap imageset=mos1_1-2keV.fits attitudeset=attitude.fits eventset=mos1_filt_time.fits
expimageset=mos1_expmap_1-2keV.fits pimin=1000 pimax=2000
Extract the Source and Background Spectra
Throughout the following, please keep in mind that some parameters are instrument-dependent. The parameter specchannelmax should be set to 11999 for the MOS, or 20479 for the PN. Also, for the PN, the most stringent filters, (FLAG==0)&&(PATTERN<=4), must be included in the expression to get a high-quality spectrum.For the MOS, the standard filters should be appropriate for many cases, though there are some instances where tightening the selection requirements might be needed. For example, if obtaining the best possible spectral resolution is critical to your work, and the corresponding loss of counts is not important, only the single pixel events should be selected (PATTERN==0). If your observation is of a bright source, you again might want to select only the single pixel events to mitigate pile up.
In any case, you'll need to know spatial information about the area over which you want to extract the spectrum; this can be obtained by downloading the filtered event file and displaying it on your local machine with ds9. Select the object whose spectrum you wish to extract and resize as desired. For this example, we will choose the source at (26188.5,22816.5) and set the extraction radius to 300 (in physical units).
Then, to extract the source spectrum,
evselect table='mos1_filt_time.fits' energycolumn='PI' filteredset='mos1_filtered.fits'
filtertype='expression' expression='((X,Y) in CIRCLE(26188.5,22816.5,300))'
spectrumset='mos1_pi.fits' spectralbinsize=5
withspecranges=yes specchannelmin=0 specchannelmax=11999
where the parameters are as indicated previously,
and
-
spectrumset - name of output spectrum
spectralbinsize - size of bin, in eV
withspecranges - covering a certain spectral range
specchannelmin - minimum of spectral range
specchannelmax - maximum of spectral range
evselect table=mos1_filt_time.fits energycolumn='PI' filteredset='bkg_filtered.fits'
filtertype='expression' spectrumset='bkg_pi.fits' spectralbinsize=5
expression='((X,Y) in CIRCLE(26188.5,22816.5,1500))&&!((X,Y) in CIRCLE(26188.5,22816.5,500))'
withspecranges=yes specchannelmin=0 specchannelmax=11999
Determine the Spectrum Extraction Areas
The source and background region areas can now be found. This is done with the task backscale, which takes into account any bad pixels or chip gaps, and writes the result into the BACKSCAL keyword of the spectrum table. To find the source and background extraction areas:
backscale spectrumset=mos1_pi.fits badpixlocation=mos1_filt_time.fits
backscale spectrumset=bkg_pi.fits badpixlocation=mos1_filt_time.fits
Check for Pile Up
Depending on how bright the source is and what modes the EPIC detectors are in, event pile up may be a problem. Pile up occurs when a source is so bright that incoming X-rays strike two neighboring pixels or the same pixel in the CCD more than once in a read-out cycle. In such cases the energies of the two events are in effect added together to form one event. If this happens sufficiently often, 1) the spectrum will appear to be harder than it actually is, and 2) the count rate will be underestimated, since multiple events will be undercounte d. To check whether pile up may be a problem, use the SAS task epatplot. (Heavily piled sources will be immediately obvious, as they will have a "hole" in the center.) Note that this procedure requires as input the event files created when the spectrum was made.
To check for pile up in our Lockman Hole example,
epatplot set=mos1_filtered.fits plotfile=mos1_epat.ps useplotfile=yes
withbackgroundset=yes backgroundset=bkg_filtered.fits
where
-
set - name of the event file
useplotfile - use file name indicated in plotfile for output?
plotfile - output file name
withbackgroundset - use the background event set for background subtraction?
backgroundset - name of background event file
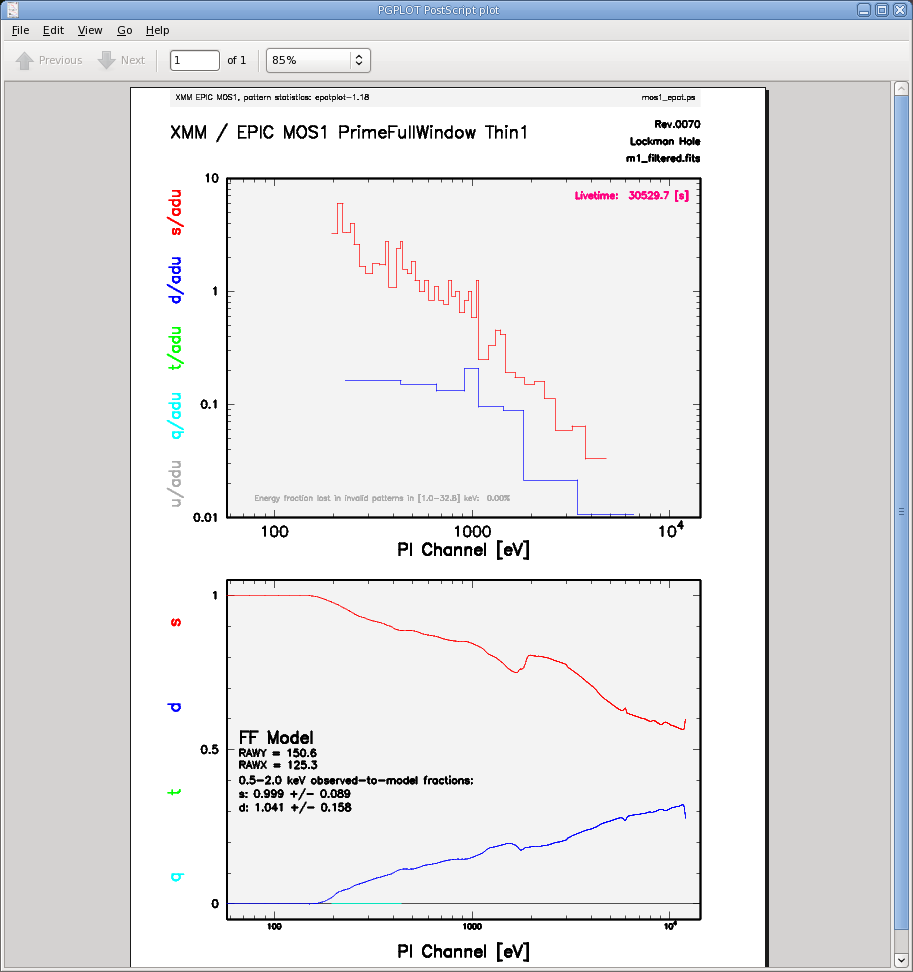
The output of epatplot is a postscript file, mos1_epat.ps, which may be downloaded and viewed with gv, containing two graphs describing the distribution of counts as a function of PI channel, as seen in Figure 4 (left).
A few words about interpretting the plots are in order. The top is the distribution of counts versus PI channel for each pattern class (single, double, triple, quadruple), and the bottom is the expected pattern distribution (smooth lines) plotted over the observed distribution (histogram). The lower plot shows the model distributions for single and double events and the observed distributions. It also gives the ratio of observed-to-modeled events with 1-σ uncertainties for single and double pattern events over a given energy range. (The default is 0.5-2.0 keV; this can be changed with the pileupnumberenergyrange parameter.) If the data is not piled up, there will be good agreement between the modeled and observed single and double event pattern distributions. Also, the observed-to-modeled fractions for both singles and doubles in the 0.5-2.0 keV range will be unity, within errors. In contrast, if the data is piled up, there will be clear divergence between the modeled and observed pattern distributions, and the observed-to-modeled fraction for singles will be less than 1.0, and for doubles, it will be greater than 1.0.
Finally, when examining the plots, it should noted that the observed-to-modeled fractions can be inaccurate. Therefore, the agreement between the modeled and observed single and double event pattern distributions should be the main factor in determining if an observation is affected by pile up or not.
The source used in our Lockman Hole example is too faint to provide reasonable statistics for epatplot and is far from being affected by pile up. For comparison, an example of a bright source (from a different observation) which is strongly affected by pileup is shown in Figure 4 (right). Note that the observed-to-model fraction for doubles is over 1.0, and there is severe divergence between the model and the observed pattern distribution.

|
My Observation is Piled Up! Now What?
If you're working with a different (much brighter) dataset that does show signs of pile up, there are a few ways to deal with it. First, using the region selection and event file filtering procedures demonstrated in earlier sections, you can excise the inner-most regions of a source (as they are the most heavily piled up), re-extract the spectrum, and continue your analysis on the excised event file. For this procedure, it is recommended that you take an iterative approach: remove an inner region, extract a spectrum, check with epatplot, and repeat, each time removing a slightly larger region, until the model and observed distribution functions agree. You can also use the event file filtering procedures to include only single pixel events (PATTERN==0), as these events are less sensitive to pile up than other patterns.
Create the Photon Redistribution Matrix (RMF) and Ancillary File (ARF)
Now that a source spectrum has been extracted, we need to reformat the detector response by making a redistribution matrix file (RMF) and ancillary response file (ARF). To make the RMF:
rmfgen rmfset=mos1_rmf.fits spectrumset=mos1_pi.fits
where
-
rmfset - output file
spectrumset - spectrum file
Now use the RMF, spectrum, and event file to make the ancillary file.
arfgen arfset=mos1_arf.fits spectrumset=mos1_pi.fits withrmfset=yes
rmfset=mos1_rmf.fits withbadpixcorr=yes badpixlocation=mos1_filt_time.fits
where
-
arfset - output ARF file name
spectrumset - input spectrum file name
withrmfset - flag to use the RMF
rmfset - RMF file created by rmfgen
withbadpixcorr - flag to include the bad pixel correction
badpixlocation - file containing the bad pixel information; should be set to the event
file from which the spectrum was extracted.
The spectrum can be fit using HEASoft or CIAO packages, as SAS does not include fitting software.
If you have any questions concerning XMM-Newton send e-mail to xmmhelp@lists.nasa.gov