THE XMM-NEWTON ABC GUIDE, STREAMLINED
RGS, Hera Command Window
Contents
Prepare the Data
Reprocess the Data
Make Images
Make a Light Curve
Generate and Apply a New GTI
Check for Pile Up
Create the Photon Redistribution Matrix (RMF)
Combining Spectra
Prepare the Data
Please note that the two tasks in this section (cifbuild and odfingest) must be run in the ODF directory. These are the only tasks with that requirement, and after this section, we will work exclusively in our reprocessing directory.Many SAS tasks require calibration information from the Calibration Access Layer (CAL). Relevant files are accessed from the set of Current Calibration File (CCF) data using a CCF Index File (CIF). Setting the environment paraters follows the same syntax as the cshell in linux. To set the environment parameters and make the ccf.cif file, navigate into the ODF directory and in the Command Window, type
setenv SAS_ODF /data/0134720401/ODF setenv SAS_ODFPATH /data/0134720401/ODF cifbuildTo use the updated CIF file in further processing, you will need to reset the environment variable SAS_CCF:
setenv SAS_CCF /data/0134720401/ODF/ccf.cifThe task odfingest extends the Observation Data File (ODF) summary file with data extracted from the instrument housekeeping data files and the calibration database. It is only necessary to run it once on any dataset, and will cause problems if it is run a second time. If for some reason odfingest must be rerun, you must first delete the earlier file it produced. This file largely follows the standard XMM naming convention, but has SUM.SAS appended to it. To run odfingest and reset the environment variable:
odfingest setenv SAS_ODF /data/0134720401/ODF/0517_0134720401_SCX00000SUM.SAS
Reprocess the Data
To reprocess the data, go up one directory in the tree and make a new working directory, either using the buttons in the upper left corner of the User Account Window or the commands you would expect with a linux machine, as seen below. When you are in the working directory, call rgsproc:cd .. mkdir reproc cd reproc rgsprocThis takes several minutes, and outputs 12 files per RGS, plus 3 general use FITS files.
A word about renaming files in Hera: files cannot currently be copied, only renamed. However, renaming can have unexpected ramifications when processing RGS files. Therefore, it is generally recommended at present that the user NOT RENAME RGS FILES from their default rgsproc output names. Users are free to name the output from all other tasks as they see fit.
The pipeline task rgsproc is very flexible and can address potential pitfalls for RGS users. If the default parameters are sufficient for your data (and they should be for most), feel free to skip ahead. However, there are some cases where the standard processing may not be appropriate. These are discussed next.
First, if the data includes a nearby bright optical source, with certain pointing angles, zeroth-order optical light may be reflected off the telescope optics and onto the RGS CCD detectors. If this falls on an extraction region, the current energy calibration will require a wavelength-dependent zero-offset. Stray light can be detected on RGS DIAGNOSTIC images taken before, during and after the observation. This test, and the offset correction, are not performed on the data before delivery. To check for stray light and apply the appropriate offsets,
rgsproc calcoffsets=yes withoffsethistogram=nowhere
-
calcoffsets - calculate PHA offsets from diagnostic images
withoffsethistogram - produce a histogram of uncalibrated excess for the user
If you are not using this example data set of Capella, your data might include a nearby bright X-ray background source that is well-separated from the target in the cross-dispersion direction. A mask can be created that excludes it from the background region. This can be done by identifiying the source in the EPIC images, and taking its coordinates from the EPIC source list which is included among the pipeline products. For example, lets say that the bright neighboring object is found to be the third source listed in the sources file, and the first source is the target:
rgsproc orders='1 2' withepicset=yes epicset=PiiiiiijjkkaablllEMSRLInmmm.FTZ
exclsrcsexpr='INDEX==1&&INDEX==3'
where
-
withepicset - calculate extraction regions for the sources contained in
an EPIC source list
epicset - name of the EPIC source list, such as generated by emldetect
or eboxdetect procedures
exclsrcsexpr - expression to identify which source(s) should be excluded
from the background extraction region
If the true coordinates of an object are not included in the EPIC source list or the science proposal, the user can define the coordinates of a new source:
rgsproc withsrc=yes srclabel='Capella' srcstyle=radec srcra=79.1708 srcdec=+45.9981where
-
withsrc - make the source be user-defined
srclabel - source name
srcstyle - coordinate system in which the source position is defined
srcra - the source's right ascension in decimal degrees
srcdec - the source's declination in decimal degrees
Make Images
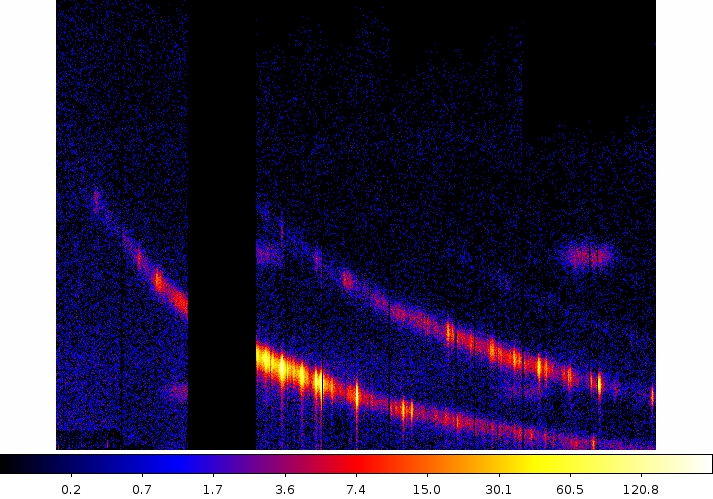
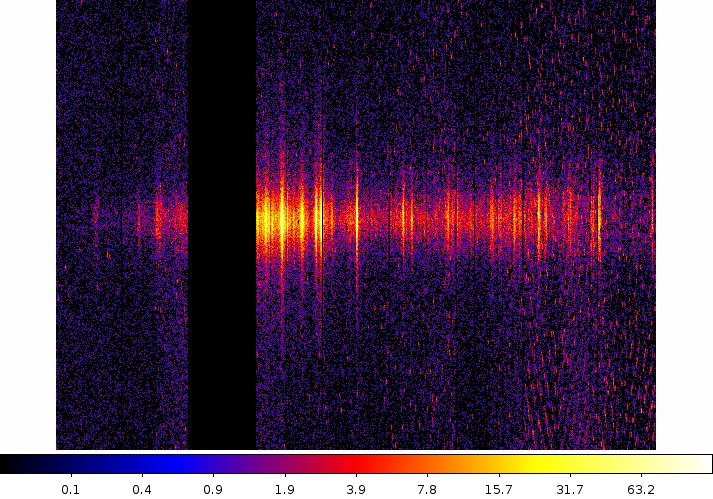
Two commonly-made plots are those showing PI vs. BETA_CORR (also known as "banana plots") and XDSP_CORR vs. BETA_CORR.To make a banana plot, type
evselect table=P0134720401R1S007EVENLI0000.FIT imageset=pi_bc.fits xcolumn=BETA_CORR ycolumn=PI
imagebinning=imageSize ximagesize=600 yimagesize=600
where
-
table - make the source be user-defined
imageset - source name
xcolumn - coordinate system in which the source position is defined
ycolumn - the source's right ascension in decimal degrees
imagebinning - the source's declination in decimal degrees
ximagesize - size of the image on x axis
yimagesize - size of the image on y axis
|
Make a Light Curve
The XMM-Newton Observatory is susceptible to soft particle flaring, so it is necessary to examine the light curve to determine how much of the data is useful. We will extract a region, CCD9, that is most susceptible to these events and generally records the least source events due to its location close to the optical axis. Also, to avoid confusing background for source variability, a region filter that removes the source from the final event list should be used. The region filters are kept in the source file product P*SRCLI_*.FIT.
To create a light curve, type
evselect table=P0134720401R1S007EVENLI0000.FIT rateset=r1_ltcrv.fits maketimecolumn=yes
timebinsize=100 makeratecolumn=yes
expression=
'(CCDNR==9)&&(REGION(P0134720401R1S007SRCLI_0000.FIT:RGS1_BACKGROUND,M_LAMBDA,XDSP_CORR))'
where
-
table - input event table
rateset - name of output light curve file
maketimecolumn - make a time column
timebinsize - time binning (seconds)
makeratecolumn - make a count rate column, otherwise a count column will be created
expression - filtering criteria
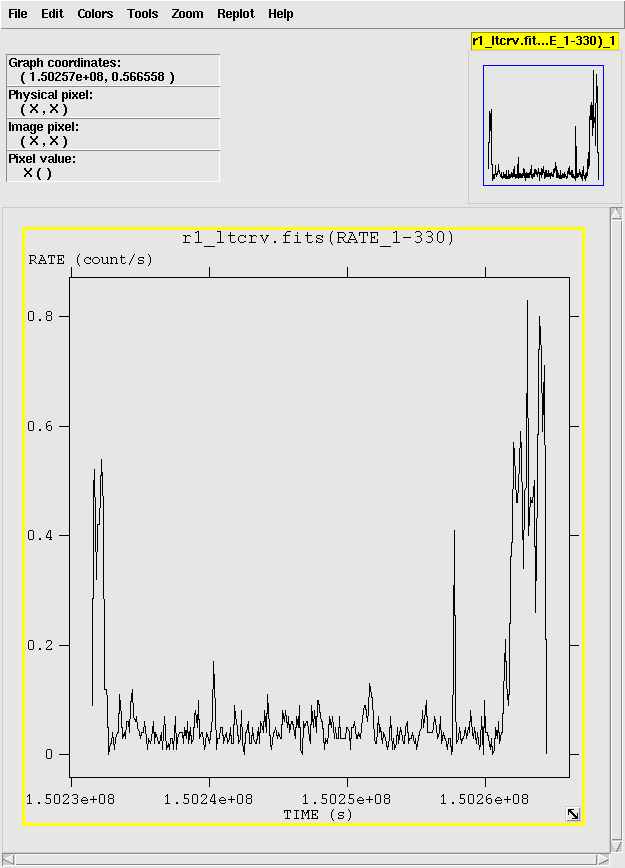
The output file r1_ltcrv.fits can be download to the user's machine and viewed
with fv, as shown in Figure 2.
- fv r1_ltcrv.fits &
Generate and Apply a New GTI
Examination of the lightcurve shows that there are high flux times at the start and end of the observations. Lets say we're interested in the quiescent periods. To extract them, we need to make an additional Good Time Interval (GTI) file and apply it by rerunning rgsproc.
There are two tasks that make a GTI file: gtibuild and tabgtigen. Either will produce the needed file, so which one to use is a matter of the user's preference. Both are demonstrated below.
Filter on TIME With gtibuild
The first method, using gtibuild, requires a text file as input. The file must be made on the user's local machine and uploaded to Hera. In the first two columns, enter the start and end times (in seconds) that you are interested in, and in the third column, indicate with either a + or - sign whether that region should be kept or removed. Each good (or bad) time interval should get its own line. In the example case, we would write in our text file (named gti.txt):
1.50233e8 1.50258e8 +Then, we upload it to Hera and proceed to gtibuild:
gtibuild file=gti.txt table=gti.fitswhere
-
file - input text file
table - output GTI table
Filter on TIME With tabgtigen
Alternatively, we could make a filter on TIME using tabgtigen using the filtering expression from the good times noted previously:
tabgtigen table=r1_ltcrv.fits gtiset=gtiset.fits timecolumn=TIME \
expression='(TIME in [1.50233e8:1.50258e8])'
where
-
table - input data set (the light curve)
gtiset - output GTI file name
timecolumn - name of column that contains the time stamps
expression - filtering expression
Filter on RATE With tabgtigen
Finally, we could make a filter on RATE using tabgtigen. The typical quiescent count rate for this observation is about 0.1 ct/s, so we would have:
tabgtigen table=r1_ltcrv.fits gtiset=gtiset.fits timecolumn=TIME \
expression='(RATE <= 0.1)'
where parameters are as described above.
Apply the New GTI File
We can apply the GTI to the event file by running rgsproc again. rgsproc is a complex task, running several steps, with five different entry and exit points. It is not necessary to rerun all the steps in the procudure, only the ones involving filtering.
rgsproc auxgtitables=gti.fits entrystage=3:filter finalstage=5:fluxingwhere
-
auxgtitables - gti file in FITS format
entrystage - stage at which to begin processing
finalstage - stage at which to end processing
Check for Pile Up
Depending on how bright the source is event pile up may be a problem. Pile up occurs when a source is so bright that incoming X-rays strike two neighboring pixels or the same pixel in the CCD more than once in a read-out cycle. In the RGS, it will cause two coincident 1st order photons to combine into a single 2nd order event at the same spatial position on the detector but at half the wavelength; this results in photons moving from 1st order to 2nd order, and 2nd order to 3rd. Further, it can increase the number of complicated event patterns of the sort that the on-board processing removes; this results in differences in pile up between the two RGS detectors, and between CCD locations.
It is always a good idea to check if your observation is piled up, especially if the object is bright (total count rate in all orders > 12 cts/s in RGS1, and 6 cts/s in RGS2).
One way to determine if there is pile up is to examine the ratio of 1st and 2nd order fluxed spectra. Non-piled sources should have fluxed spectra that agree within a few percent, while piled sources will show discrepancies of about 10% or higher.
When rgsproc is run through the 5:fluxing stage, one of the outputs is fluxed spectra. They follow the naming convention *OBX*fluxed1000.FIT and *OBX*fluxed1000.FIT, for the 1st and 2nd orders, respectively. They can be viewed by downloading them and opening them up with fv.
If for some reason rgsproc wasn't run through that stage, fluxed spectra are easy enough to produce with rgsfluxer. Please note that rgsfluxer requires as input the response files (RMFs) for the spectra; these are produced by rgsproc (and have the nomenclature *RSPMAT*.FIT) or, alternatively, can be generated with the task rgsrmfgen as discussed below. Each RGS instrument and each order will have its own RMF.
To make fluxed order 1 spectra,
rgsfluxer pha='P0134720401R1S007SRSPEC1001.FIT P0134720401R2S008SRSPEC1001.FIT' \
rmf='P0134720401R1S007RSPMAT1001.FIT P0134720401R2S008RSPMAT1001.FIT' \
file=fluxed_o1.fits
where
-
pha - list of spectrum files
rmf - list of response matrices
file - output spectrum
Fluxed 2nd order spectra are made in a similar way.
Unfortunately, if your spectrum is piled up, there is not a lot you can do to fix it; there is no tried-and-true method like with the EPIC and what there is applies only to certain cases, such as if you assume the 2nd order spectrum is composed entirely of piled up 1st order photons. For more information, see Ness et al. 2007, ApJ, 665, 1334.
Create the Photon Redistribution Matrix (RMF)
The task rgsproc generates a response matrix automatically (with the nomenclature *RSPMAT*.FIT), but as noted above the source coordinates are under the observer's control. The source coordinates have a profound influence on the accuracy of the wavelength scale as recorded in the RMF that is produced automatically by rgsproc, and each RGS instrument and each order will have its own RMF.
Making the RMF is easily done with rgsrmfgen. Please note that, unlike with EPIC data, it is not necessary to make ancillary response files (ARFS).
To make the RMFs for the first order RGS1 spectrum,
rgsrmfgen spectrumset=P0134720401R1S007SRSPEC1001.FIT rmfset=r1_o1_rmf.fits
evlist=P0134720401R1S007EVENLI0000.FIT emin=0.4 emax=2.5 rows=5000
where
-
spectrumset - spectrum file
evlist - event file
emin - lower energy limit of the response file
emax - upper energy limit of the response file
rows - number of energy bins; this should be greater than 3000
rmfset - output FITS file
At this point, the spectra can be either analyzed or combined with other spectra. However, if your source is very bright (total count rate in all orders > 12 cts/s in RGS1, and 6 cts/s in RGS2), it is a good idea to check it for pile up first.
Combining Spectra
Spectra from the same order in RGS1 and RGS2 can be safely combined to create a spectrum with higher signal-to-noise if they were reprocessed using rgsproc with spectrumbinning=lambda, which is the default setting. The task we will use to merge source spectra, rgscombine also merges response files and background spectra. When merging response files, be sure that they have the same number of bins. For this example, we assume that RMFs were made for order 1 in both RGS1 and RGS2 with rgsproc, which has a default bin number of 4000.To merge RGS1 and RGS2 spectra,
rgscombine pha='P0134720401R1S007SRSPEC1001.FIT P0134720401R2S008SRSPEC1001.FIT'
rmf='P0134720401R1S007RSPMAT1001.FIT P0134720401R2S008RSPMAT1001.FIT'
bkg='P0134720401R1S007BGSPEC1001.FIT P0134720401R2S008BGSPEC1001.FIT'
filepha='r12_o1_srspec.fits' filermf='r12_o1_rmf.fits'
filebkg='r12_o1_bgspec.fits' rmfgrid=4000
where
-
pha - list of spectrum files
rmf - list of response matrices
bkg - list of bakcground spectrum files
filepha - output merged spectrum
filermf - output merged response matrix
filebkg - output merged badkground spectrum
rmfgrid - number of energy bins; should be the same as the input RMFs
If you have any questions concerning XMM-Newton send e-mail to xmmhelp@lists.nasa.gov