Come analyze HEASARC, IRSA, and MAST data in the cloud! The Fornax Initiative is now welcoming all interested beta users.
HEASARC@SciServer User Guide
A list of the latest software version available on SciServer can be found in the build.json on GitHub.
1. Getting started

- Create an account at https://sciserver.org
- Go to the Compute page from the home page or by selecting it from the grid icon at top right of the page.
- Create a container, select the HEASARC compute image (e.g. HEASARCv6.35.1), select a user volume, and finally make sure HEASARC data is checked under data volumes.
- Go to the container by clicking its name in the list of containers. This will launch the JupyterLab interface, which gives you access to a virtual machine with HEASoft and other high energy software installed, along with all the HEASARC data available locally. You can either work on a notebook (python) or launch a terminal.
2. Containers
The SciServer containers managed by the HEASARC are labeled by the HEASoft version (e.g. HEASARCv6.35.1), and contain other software as well (e.g. ciao, XMM SAS etc.). The HEASARC also maintains the NAVO-workshop container image, which contains tutorials on how to use python to discover and access NASA data using the virtual observatory. See Data Discovery for details.
You can create up to 3 containers at the same time. You can start, stop and delete the containers from the Containers page.
Containers not in use will be stopped automatically. You can restart them by clicking the green start icon.
Closing the browser when a container is running does not stop it.
A running container is accessed through a Jupyter Lab interface. The different software is accessed through different conda environments (e.g. heasoft, ciao, xmmsas etc.). See the software section below for details.
To access the archive or your user data within the container, the relevant volumes need to be selected when creating the container.
When you create a new container, you get a user and machine name like: idies@f0b78ddd5226
The username idies is the same for everyone. The machine name is a unique hash for that container.
3. File systems
There are two types of storage areas: Temporary and Persistent.
- The content of the persistent area persists between sessions, and can be accessed from multiple containers. It has a limit of 10GB.
- The temporary area has larger disk space, but its content is not guarenteed to persist between sessions.
These storage areas are available under:
/home/idies/workspace/Storage/<username>/persistent/and
/home/idies/workspace/Temporary/<username>/scratch/
If you define your own user volume (for sharing with another user, see below) and mount it to the container, it will also appear under Storage, as will any other volume that is shared with you by other users under their username:
/home/idies/workspace/Storage/<username>/<my_user_volume>/
/home/idies/workspace/Storage/<anotherUser>/<their_user_volume>/
See Miscellaneous below for backup discussion.
When you mount the HEASARC data volume when creating the container, it will be available under:
/home/idies/workspace/headata/FTP/
or
/FTP/
The FTP area will contain all of the HEASARC data holdings exactly as they are organized in the main HEASARC website.
The default software installation is not on that volume but built within the compute container under
/opt/heasoft
Your initial home directory is always /home/idies. That home directory is created new from the image with each new container and is not part of your persistent storage. Anything stored in that homedir (such as edits to a ~/.xspec/Xspec.init file or your ~/.bashrc file), it will not persist to a new container.
4. Files, groups and sharing
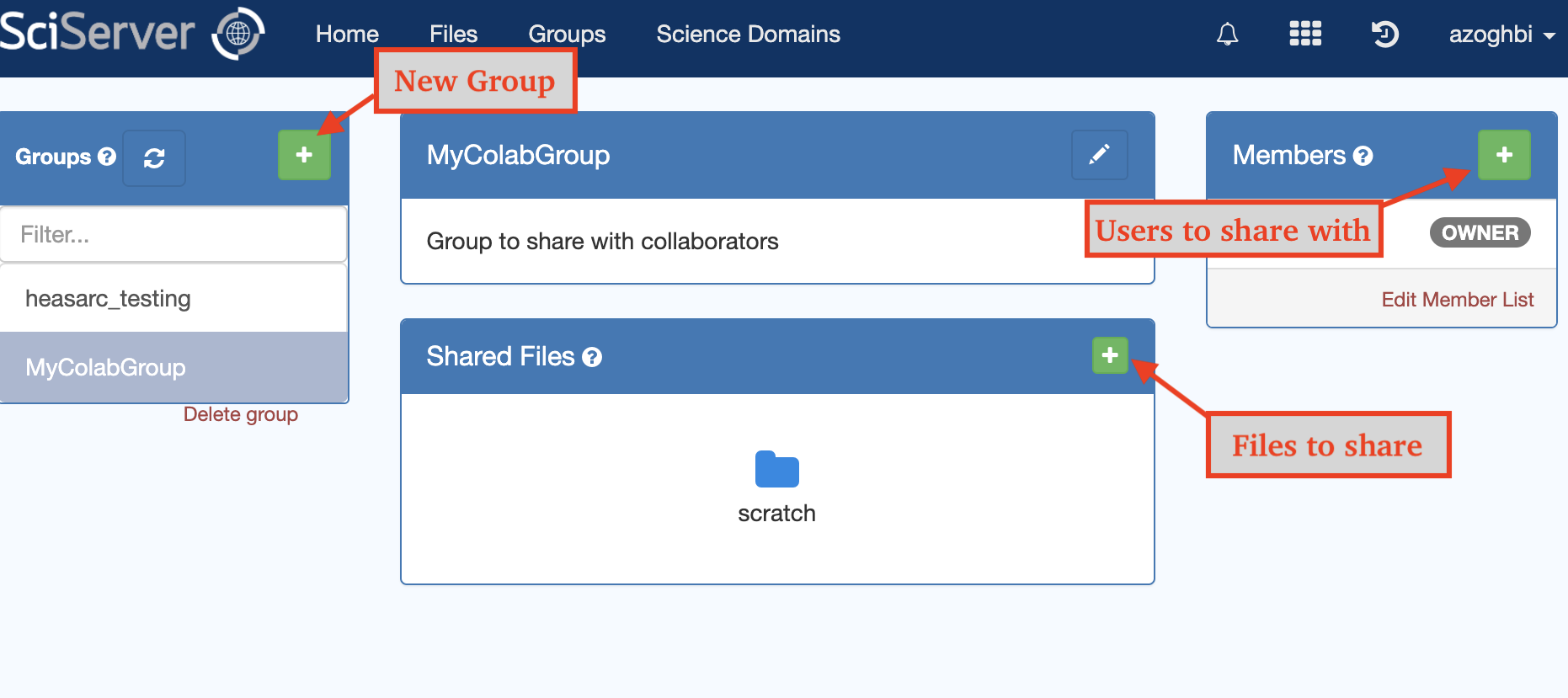
The Files app (as apposed to the Compute app discussed above), offers the option to browse the data volumnes, including the user, shared and data volumes. There, you can download and upload files.
The Groups app is where groups and sharing is managed. You can create your own groups, add your own shared files/volumes, and invite your collaborators to join your group. This way, you and your group can share data that are not available to anybody else.

File download and upload to the persistent of temporary space can be done either from the Files app, or from the Jupyter Lab file browser when the container is running (right-click to download and upload with button on top). You can also use other tools like wget, curl, git or scp inside the container to download and/or upload files to/from the internet.
5. Software
The HEASARC compute image has several software environments, which are managed through conda. The default environment is heasoft, which contains the full HEASoft installation.
In the terminal, the prompt will contain the name of the environment. It may look something like:
(heasoft) idies@9dbc1849c28c:~$
This means that the heasoft environment is active, and one can run all the standard HEASoft tools, for example:
(heasoft) idies@9dbc1849c28c:~$ fversion 21Mar2024_V6.33.1 (heasoft) idies@9dbc1849c28c:~$ which fversion /opt/heasoft/x86_64-pc-linux-gnu-libc2.35/bin/fversion
The calibration database CALDB is set to the archive's calibration area, which will be kept up to date:
$ echo $CALDB /home/idies/workspace/headata/FTP/caldb
Additional environments include: ciao, xmmsas, fermi, spex. For example, to activate the ciao environment:
(heasoft) idies@9dbc1849c28c:~$ conda activate ciao (ciao) idies@9dbc1849c28c:~$
After which, all ciao tools can be run.
The same for the other environments, e.g:
(ciao) idies@9dbc1849c28c:~$ conda activate xmmsas (xmmsas) idies@9dbc1849c28c:~$ sasversion ...

When using notebooks, the environment can be selected by choosing a kernel from the list at top right part of the notebook (See image).
Additional software
If a software you need is not available, you can install it. For example, python packages can be installed with:
pip install scikit-learn pip install umap-learn
If you want to install it in a specific environment, make sure that environment is activated first.
Fortran and C/C++ compilers are also available if you want to compile your own code.
Note that such installs will persist in this container if you stop and restart it, but they will not be available in other containers.
We welcome suggestions for additional software and packages that can be added to the software set.
6. Tutorial notebook
Several example notebooks that are available once a computer container is launched. They can be found under HEASARC data, so if you mount this to your container, then you will find them at:
ls ~/workspace/sciserver_cookbooks analysis-ixpe-example.md analysis-rxte-spectra.md data-access.md ...
Note that sciserver_cookbooks folder is not in the persistent storage. Therefore, any changes you make to these notebooks will not be saved. Move the notebooks to the persistent storage area if you want your changes to be saved. You can do this from either the terminal or from the Jupyter file browser.
The tutorial folder (also available on the HEASARC GitHub page) contains:
- The introduction notebook gives an overview of the tutorial content.
- Notebooks whose name start with data-... contain tutorials on finding and accessing data.
- Notebooks that start with analysis-... give example of analysing data from different missions in the heasoft environment.
- Examples for analysing XMM data are under the xmm folder.
- Notebooks covering other topics (e.g. image analysis with jdaviz or JS9) start with misc-...
7. Data discovery with HEASARC tools
The usual ways of discovering HEASARC data with our Browse and Xamin tools through the website. For Xamin, when you have a list of files or observations, there an option to generate a list of SciServer's paths (starting with /FTP/) that you can then copy-paste into SciServer as needed. Those paths will work on SciServer within your container when you use the HEASARC image and mount the HEASARC data volume. For W3Browse, you can get a download script and then edit that script into a simple file list starting /FTP.
The data archive can also be search and explored using the Virtual Observatory (VO) python interface. You can see example of doing this using the python package pyvo in the notebook tutorials whose names start with data-....
More detailed examples of using the VO's powerful python interface are available in the NASA Virtual Observatory notebooks tutorials. You can either download them during your compute session, or use the NAVO-workshop image when creating the compute container. That image has the system setup to run the tutorial notebooks (Note that HEASoft is not available in that image).
8. Batch
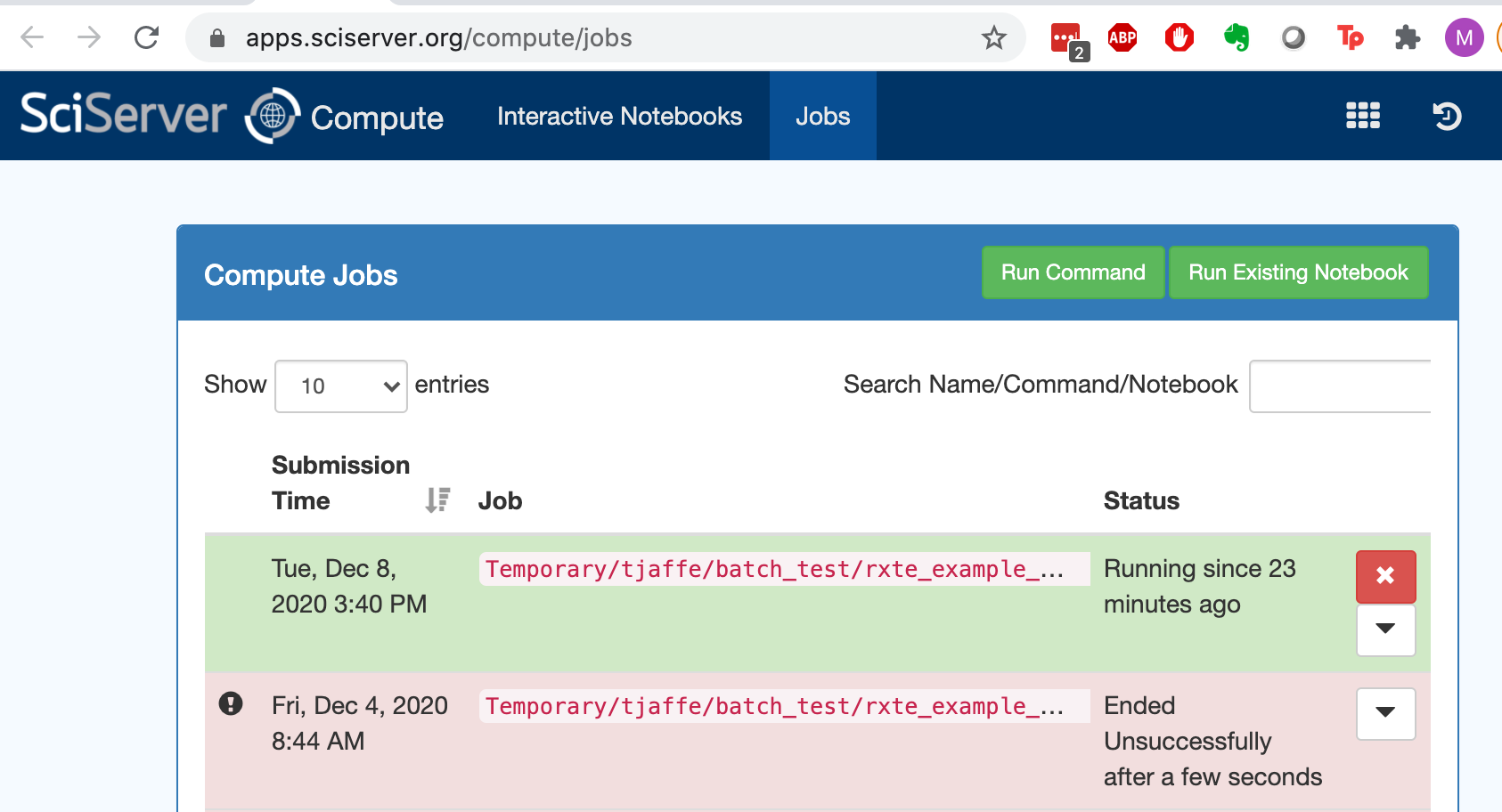
You can submit a notebook or a script that you have tested interactively to the batch for processing. The batch service is called Compute Jobs (while Compute is interactive) under the main menus. Note that you need to set up your Jupyter notebook to use the correct (e.g. heasoft) kernel before submitting it to the batch. This is done by ensuring the correct kernel is selected when the notebooks is open, and then saved so that the name of the kernel is also saved in the notebook metadata
Submitting to the batch starts with a process a bit like setting up a new container. You have to select the compute image and the volumes that you want to mount. If it starts successfully, it will look something like this:
9. Image Analysis
Many high energy pipelines will require some type of image analysis, from simple source and image creation to complex analysis. The standard tools for such analysis is DS9. Unfortunately, DS9 is a desktop application and not optimized for running in a platform like SciServer over a browser. Currently, DS9 is available on SciServer through an Xserver. You can start it by clicking the DS9 icon in the JupyterLab laucher. Note that is launches a full desktop server, and therfore can be slow for interactive analysis. See next for alternatives. Once an Xserver is launched (by launching DS9), other applications can use it too, for example, you can start an xspec session in the terminal, and set the plotting device to xwindow, it will show up along with the DS9 applications. The same applies to tools such as FV and Ximage.
The recommeded alterntive to DS9 is JS9, which has many of the features available in DS9. In addition to being used for interactive analysis, JS9 can be used inside a jupyter notebook and included as part of the analysis flow. An example showing how to use JS9 is given in the misc-js9-demo notebook.
The notebook misc-jdaviz-demo provides another way of doing image analysis inside a notebook using jdaviz. This is the image analysis tool developed for JWST which` has many applications.
10. Notes
Citing
To cite the use of HEASARC on SciServer, please use these citations:
If HEASoft and FTOOLS have been useful, please use the citation at the bottom of the FTOOLS page.
Help
If you encounter any issues running code or accessing the data, please contact the HEASARC help desk. If your issue is not related to the HEASARC setup specifically, then you can email the SciServer helpdesk. Click on your username at the top right of your dashboard and select Help from the dropdown menu.
If you could use some additional software that you cannot install yourself, you can also ask at the HEASARC help desk if we can install it for you.
Limits
A summary of users' limitations on SciServer:
- Disk space (persistent): 10 GB total, including your 'persistent' area and all of your user volumes (not including user volumes shared with you).
- Disk space (temporary): unlimited for 72 hours from file creation; not private.
- Number of containers: you may only have three containers defined at a time (whether running or not).
- Lifetime of containers: 90 days beyond last access; i.e., if unused for 90 days, they may be removed. Large Jobs: 32 CPU, 240GB RAM. Run for 16 hours. Small Jobs: 32 CPU, up to 6 jobs, with 32GB/job. Run for 1 hour each.
- Batch (asynchronous) time: HEASARC users are currently limited to the "Small Jobs Domain" of 1 hour with access to 32 CPU, up to 6 jobs, with 32GB/job.
- Large batch: 32 CPU, 240GB RAM and run for a maximum of 16 hours.
See also https://www.sciserver.org/support/policies/.
User Contributions
One of the benefits of SciServer is how easy it makes to share data, code, results, etc. among collaborators. But you can also contribute them to the community of HEASARC@SciServer users. Please reach out to the HEASARC help desk if you have suggestions or comments. We also have a notebooks repository on GitHub to which you can contribute through submitting issues to request changes, pull requests to contribute, etc. Likewise for the PyXspec Jupyter notebooks repo.
Miscellaneous
In our experience, if you leave the browser window open a long time, SciServer stays connected, though it starts to pop up little windows with apparently minor access errors. But if you start it on VPN and the VPN connection closes, then you lose your access to it. (Note that your running containers are still running, your running notebooks are in the same state you left them, you just have to reconnect to the Jupyter session.)
Back up your work frequently, e.g., daily! You could crash your image, so anything in temporary is gone. Your persistent area and defined user volumes should be backed up, but don't rely on that alone. I tend to work in a Jupyter notebook, which you can easily download at the end of every day with a couple of clicks. If there are a lot of products, maybe make a tar file each day and download that. Keep your code in a repo on GitHub and commit and push changes regularly. We welcome suggestions for improving this given that it's not possible to rsync *into* SciServer, and you may not be able to connect into an institute behind a firewall (unless you set up ssh tunnels...)
It's convenient to make your own .bashrc and store it in your persistent storage area. Then when you create a new container and have the default setup, you can source your personal one as soon as you open your terminal to get your preferred setup.
Likewise, if you want an Xspec.init file to persist, you should move the ~/.xspec directory to your persistent storage:
mv /home/idies/.xspec /home/idies/workspace/Storage/myusername/persistent/And then create a symlink from the home dir in any new container.
ln -s /home/idies/workspace/Storage/myusername/persistent/.xspec /home/idies/.xspecDo this ideally before starting Xspec for the first time in the new container. Without that symlink, running Xspec for the first time in a new container will create a new /home/idies/.xspec dir without linking to your persistent one.
If you wish to change your default shell in a given container to tcsh, for example, you must add the following
line
c.NotebookApp.terminado_settings = {'shell_command':['/bin/tcsh']} to the
~/.jupyter/jupyter_notebook_config.py file. You then need to re-start your container, but once done, this setting
persists in that container. (New containers will not have this, though.)
Known Issues
- If you see this:
then close all of your SciServer windows and start from the Dashboard again. If that doesn't work, stop and restart your container.
- If you see this:
when you are trying to start a container, wait a few seconds and try refresh the page. If trying again repeatedly does not help, then try deleting the container and re-creating it.